
电子邮箱:
所在单位: 生命科学与技术学院
学历: 博士研究生毕业
办公地点:
性别: 男
联系方式:
学位: 博士
博士生导师: 是
硕士生导师: 是
学科: 生物学


杨铁林
教授,博士生导师
国家级青年人才
西安交通大学生命科学与技术学院副院长
生物医学信息与基因组学中心主任
陕西省杰出青年科学基金获得者
西安交通大学青年拔尖人才
Email:yangtielin@xjtu.edu.cn
地址:西安交通大学生命学院教二楼北213
邮政编码:710049

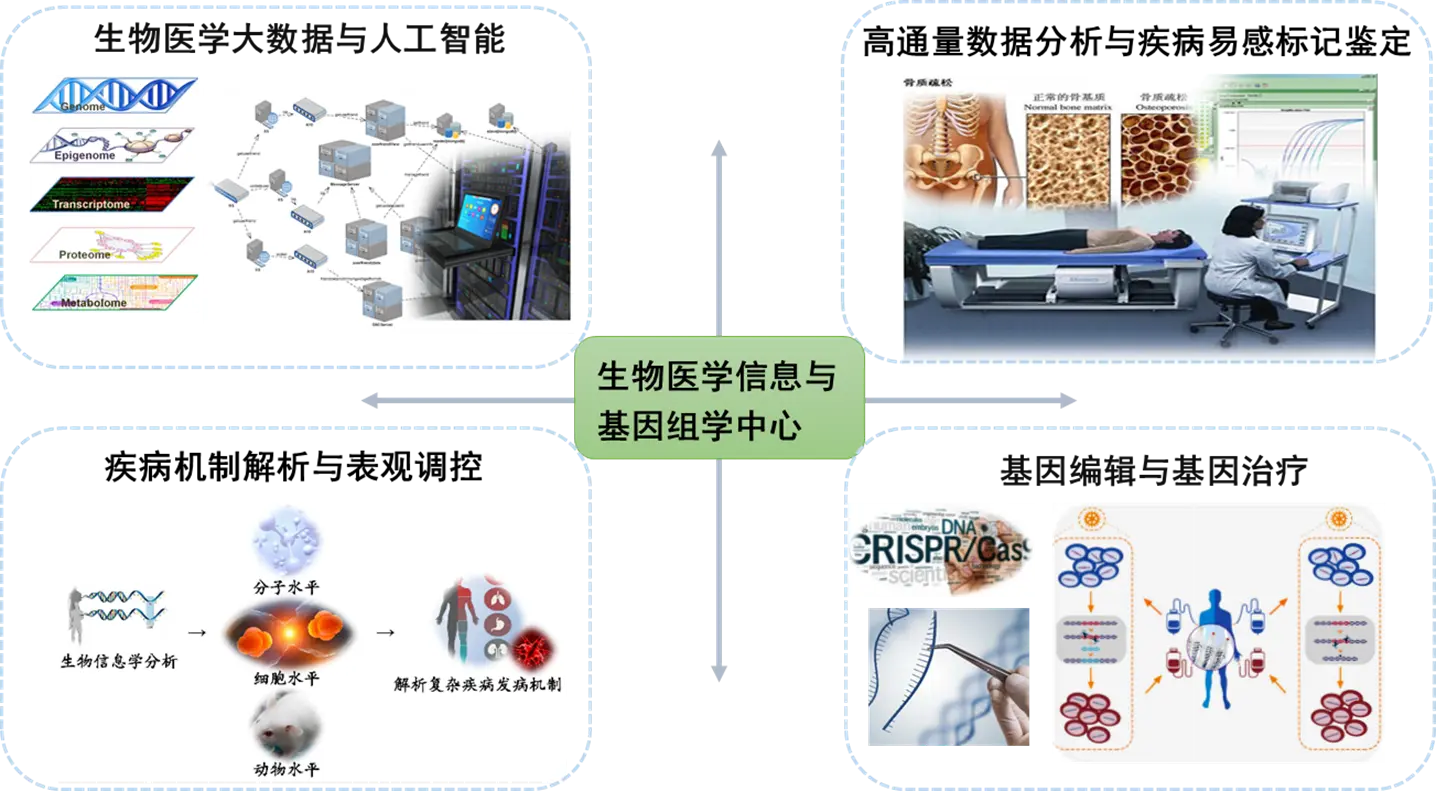
具体研究方向包括:
(1)生物医学大数据与人工智能
(2)高通量数据分析与疾病易感标记鉴定
(3)疾病机制解析与表观调控
(4)基因编辑与基因治疗
团队累计指导国家级/省级大学生创新实验30余项,本科生毕业设计40余项,其中2人获校级优秀毕业论文,10余人获院级优秀毕业论文。研究生培养方面,10余人次获研究生国家奖学金和社会奖学金,3人获校级优秀毕业论文,20余人次获“优秀毕业生”、“优秀研究生”、“优秀研究生干部”、“优秀党员”等荣誉称号。10余人次获陕西省研究生创新成果一等奖和二等奖、全国“互联网+”中国大学生创新创业大赛银奖、中国研究生未来飞行器创新大赛二等奖等奖项。中心每年派至少5名学生去美国参加国际遗传学领域顶级会议-ASHG,其中2名学生被选为学术报告,11名学生被选为优秀海报。1名博士获得国际骨质疏松症领域顶级学术会议-ESCEO-IOF的 “Young Investigator Award” 奖。2名博士生2018-2019在美国芝加哥大学和印第安纳大学联合培养,1名博士生2018参加了荷兰莱顿大学医学中心组织并资助的Eurolife summer school。中心多名博士毕业生进入西安交通大学、苏州大学等国内一流大学任教,硕士生进入华为、宝洁、广州证券、华大基因等知名企业工作。
微信公众号:生物医学信息与基因组学中心BIGC
